By Isadora Sinha
Background
All sporadic cancer is caused by acquired somatic mutations (mutations that take place within cells of the body that are not passed down/inherited). Mutations often occur by incorrect repair of DNA damage by impaired DNA pathways such as the nucleotide excision repair (NER) pathway and the DNA mismatch repair (MMR) pathway. Both these repair pathways are involved in excising the mutated nucleotide in order to repair damage to a single strand of the DNA. The pathway that is activated usually refers to the damaging agent e.g. MMR is activated when there is a mis-incorporation of bases during DNA replication and NER is activated by radiation or intercalating agents.
Whenever DNA is damaged it is up to the repair pathways to fix the damage in the best way it can. Sometimes, if the repair pathways fix it wrong, this can create a somatic mutation. Recent advances of next generation sequencing (NGS) in DNA sequencing technology have allowed for the mapping of genome-wide somatic mutations acquired in individual cancer genomes.
In this project, mutant S. cerevisiae cells deficient in one gene, msh2, involved in MMR were cultured through thousands of generations on yeast extract peptone dextrose (YPD) plates. 10 colonies were treated with UV light, a known DNA mutagen, throughout the passaging while another 10 colonies were untreated (figure 1). The resulting DNA of each colony was extracted and sequenced at passage 30 (meaning the culture was propagated onto new medium 30 times to encourage growth and reproduction). By comparing the mutations in the S. cerevisiae exposed to UV light to the “naturally” occurring somatic mutations in the untreated S. cerevisiae, the background mutations were distinguished from the mutations caused by the DNA damage due to exposure to UV light.
A clear identifiable cut-off point was determined, showing which mutations were not normal background genome instability. The results show elevated levels of mutation spread heterogeneously (distributed unevenly) through the UV treated genomes with variation in the relative contribution of the point mutations. This furthers the data found in previous studies adding to our knowledge of cancer development. This data can be compared to S. cerevisiae that have different knockouts of the NER and MMR pathways which will further our understanding of how the impaired pathways affects the repair of the DNA damaged by UV light (and other mutagens) and the impact of the resulting acquired mutations.
The mutational signatures induced can be examined by algorithms used in the cancer genome sequencing project detailed in Alexandrov, et al., 2013. This will give insight into the cause of the mutational spectra observed in individual cancer genomes, and in turn contribute to a greater understanding of how to treat individual cancers, based on their genomics.
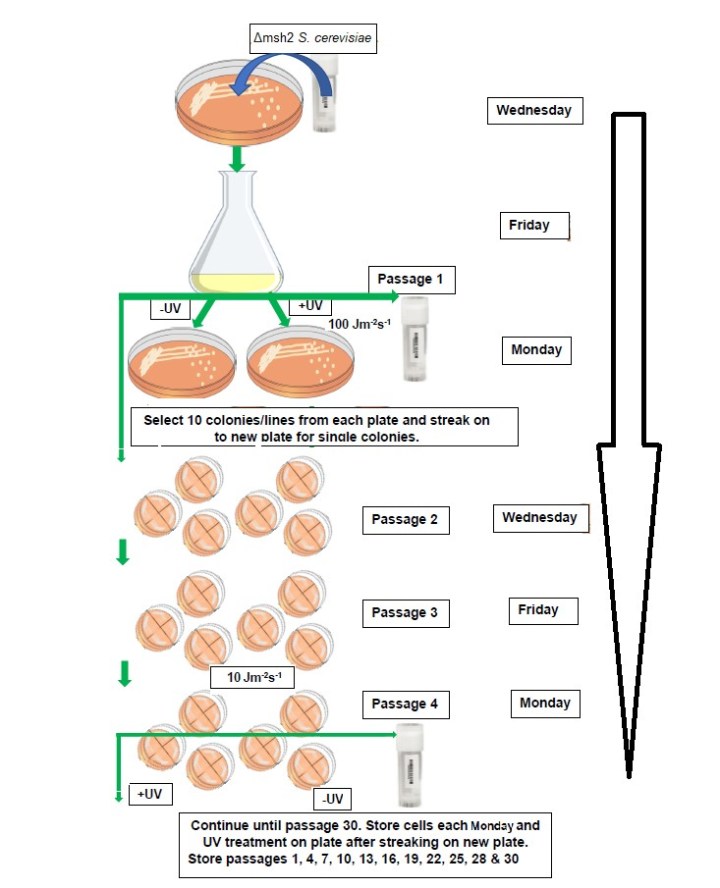
Figure 1. Outline of the yeast passaging through 30 passages with and without UV treatment.
Concluding statement
The results of this project, in addition to previous findings, indicates that the future of cancer therapy will be treating it as a chronic illness. Cancers vary so greatly genetically, even when the same type, that it is not likely there will ever be a “cure” nor a way to prevent it as genomic instability is innate and increases with age as cells undergo mitosis until the Hayflick limit (Jerry, et al., 2000) after which they either undergo apoptosis (programmed cell death) or proliferate (divide) uncontrollably (often turning into cancer). Generally, apoptosis occurs when a cell has too many mutations so it initiates and allows itself to be destroyed to keep the rest of the body safe.
Attempts to circumvent the Hayflick limit have been regarded as futile; the method of extending telomeres with telomerase in adult cells inevitability results in cancer as the telomerase appears to decrease the rate of genomic instability enabling the cell to become cancerous instead of going though cell death (Jerry, et al., 2011). Hence, the future of treating cancer seems to be by sequencing individual cancer genomes, determining the mutational signatures that led to tumorigenesis and administering the corresponding drug treatment that is compatible to the signatures found.
The cytotoxic agents used in cancer chemotherapy are mutagens themselves which often lead to secondary tumours forming that are resistant to the chemotherapy treatment the initial cancer was susceptible to (Szikriszt, et al., 2016). These secondary tumours would also have mutational histories visible within their genomes, so sequencing them would determine which chemotherapy drug is needed to treat them and the cycle would continue in an ongoing manner. In this way, cancer could be treated as a chronic condition, destroying each new cancer that develops because of previous chemotherapy treatments or inherent accumulation of somatic mutations.
References
- Alexandrov, L.B., Nik-Zainal, S., Wedge, D.C., Aparcio, S.A.J.R., Behjati, S., Biankin, A.V., Bignell, G.R., et al. (2013a) Signatures of mutational processes in human cancer. Nature. 2013 Aug 22;500(7463):415-21.
- Alexandrov, L.B., Nik-Zainal, S., Wedge, D.C., Campbell, P.J. and Stratton, M.R. (2013b) Deciphering Signatures of Mutational Processes Operative in Human Cancer. Cell Rep. 2013 Jan 31;3(1):246-59.
- Jerry W. Shay, J.W., Wright, W.E. (2000) Hayflick, his limit, and cellular ageing. Nature Reviews Molecular Cell Biology volume 1, pages 72–76.
- Jerry W. Shay, J.W., Wright, W.E. (2011) Role of telomeres and telomerase in cancer. Semin Cancer Biol. 2011 Dec; 21(6): 349–353.
- Szikriszt, B., PótiÁdám, Pipek, O., Krzystanek, M., Kanu, N., Molnár, J., Ribli, D., et al. (2016) A comprehensive survey of the mutagenic impact of common cancer cytotoxics. Genome Biology (2016) 17:99.
About me:
 I am in my final year of my BSc in Genetics at Cardiff University. I took a professional training year in the Department of Cancer and Genetics in the University Hospital of Wales followed by a summer CUROP research placement at the Sustainable Places Research Institute supervised by Dr Leanne Cullen-Unsworth. I am currently starting my final year project of detecting novel microsatellites in hedgehogs which I am excited to get started on. I have varying interests from cancer genetics to bioinformatics and conservation. My aim is to improve my knowledge and scientific acumen with further education and experience.
I am in my final year of my BSc in Genetics at Cardiff University. I took a professional training year in the Department of Cancer and Genetics in the University Hospital of Wales followed by a summer CUROP research placement at the Sustainable Places Research Institute supervised by Dr Leanne Cullen-Unsworth. I am currently starting my final year project of detecting novel microsatellites in hedgehogs which I am excited to get started on. I have varying interests from cancer genetics to bioinformatics and conservation. My aim is to improve my knowledge and scientific acumen with further education and experience.


One thought on “Investigating Somatic Mutations in Cancer Development”