By Sarah Kearns, University of Michigan, USA
There’s more to our genetics than what’s within our DNA code. Though the code itself is the basis for life, on its surface there are chemical alterations that change how it’s expressed. Because DNA’s structure is that of a helix, modifications either make it coil up or unwind where the unwound form allows for transcription. Epigenetics is the concept of chemical marks on DNA, or associated proteins that control gene activity, that act independently of the genetic code itself.
In some cases, epigenetic marks are inherited, but they can also associated with diseases or disorders such as cancers, Alzheimer’s disease and schizophrenia. Once they are in place, modifications can be passed along during cell division, a process that preserves genetic similarity across the cells in the body, but can also help propagation of disease.
Flagging and targeting disease
Epigenetic marks can be associated with diseases, but luckily they are identifiable. Too much methylation of a tumor suppressor gene will turn off its expression and propagate uncontrolled cell growth in cancer. Not enough acetyl groups and genes will be turned off leaving the cell undifferentiated – a feature of malignant cancers.
Because the relationships between epigenetic levels and disease are known, inhibitors and drug therapies have been made to target genetic or histone modifications. For example, oncogenes in most cancers have an abundance of methylation, so methylation inhibitors can reactivate anti-tumor systems. An added bonus is an inherent feature of epigenetics is reversibility because their normal biological role is to control gene expression: an off and on process. Banking off of this property, drug therapies mimic biological mechanisms and can be very effective.
Despite knowing the types of epigenetics to target, medications can become ineffective within cancer patients as there are many biological pathways that are distorted: blocking one road does not stop traffic from going around the impediment. With each individual coming to the table with their own epigenetic profile and inheritance pattern, it is difficult to know how to stop secondary disease routes without destroying vital cell functions in different pathways in the process.
An individual approach
This has lead researchers to find ways to find measurable substances within the body that indicate disease risk level, or biomarkers. Since there is already a body of data measuring modifications on a certain target gene, the goal now is to look at all the modifications across and entire genome. This individualistic approach to screening, diagnosis, and treatment is called personalized medicine. Such an approach that emphasizes the way each patient’s risk are unique and different hopes to tailor treatment combinations such that each person gets treated quickly rather than having to try a variety of different drugs while combating a disease.
Epigenomics has a lot of promise for this type of treatment because the modifications depend on overall health, on age and on environmental exposures. As such, its dynamicity and uniqueness lend themselves to a personalized approach since effective treatments for one person will be very different than another’s based on the epigenome. The tricky part is that there is not a simple test that gives information about all epigenetic marks, even though that knowledge could allow for more straightforward diagnostics and treatment plans.
Tools for diagnosis
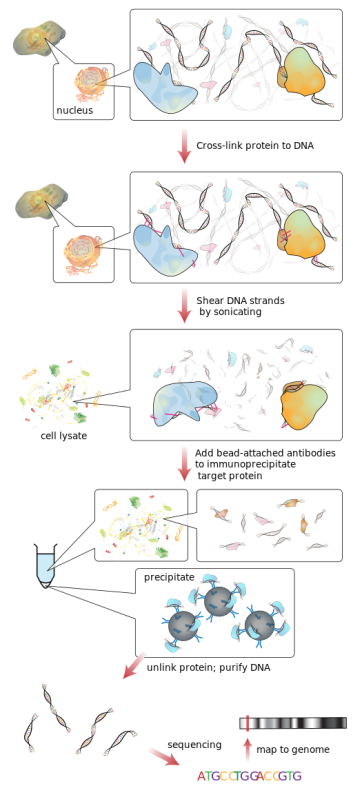
The powerful method that has been used to identify chemical modifications and their relation to disease is called ChIP, which stands for chromatin immunoprecipitation. Because chromatin in DNA coiled up around protein groups called histones, this technique is also used to look at modifications to the protein clusters as they play a big role in transcription and disease. Immunoprecipitation is a method of identifying a protein or modification of interest and pulling it out of a sample using an antibody. Just like the antibodies that find foreign agents in our bodies, specific antibodies to epigenetic modifying proteins can be engineered to flag chemical marks on chromatin.
A big drawback of traditional ChIP is that it is laborious, time consuming, and is not suitable for use at a point-of-care diagnostic laboratories. Much of this comes from needing a high number of cells, but it essentially means that this standard research method cannot be used for medicine. Work has been done to optimize, making ChIP be more flexible and usable in a clinical setting, and minimize it down to the micron level. Lab-on-a-chip technologies can take tiny (pico litre) samples and analyze them for their epigenetic content quickly drastically reducing the number of cells needed and only taking a day to obtain and analyze the data.
Future directions
These techniques will be further developed in research labs and make their way into clinical settings, figuring out ways to identify these tiny epigenetic modifications on our DNA. Having personalized maps of the chemical flags on our DNA provides more information than our genetic code alone as it gestures to whether or not our cells are in a healthy or diseased state. Knowing the combination of epigenetic marks within individualized distorted states will allow for medications to be prescribed to particular patients with higher certainty of a good outcome.
About me
 I am a current graduate student in the Chemical Biology PhD program at the University of Michigan. I’m interested in the chemical and physical properties of substrate recognition in enzymes that control epigenetic modifications to design better drug therapies. When not researching or writing about science, I am an open access advocate, an avid baker and always have at least three books on my current reading list. You can connect with me on Twitter @annotated_sci.
I am a current graduate student in the Chemical Biology PhD program at the University of Michigan. I’m interested in the chemical and physical properties of substrate recognition in enzymes that control epigenetic modifications to design better drug therapies. When not researching or writing about science, I am an open access advocate, an avid baker and always have at least three books on my current reading list. You can connect with me on Twitter @annotated_sci.
This post is the second in our epigenetics series. The first, Epigenetics: past and present by David Hornby, was published earlier this month. If you are interested in reading more on this topic, you can also check out the October issue of The Biochemist magazine on the theme of epigenetics.


2 thoughts on “Epigenetic tags for personalized diagnoses”