By Zoe Angel
CRISPR/Cas9 gene editing and treating genetic diseases
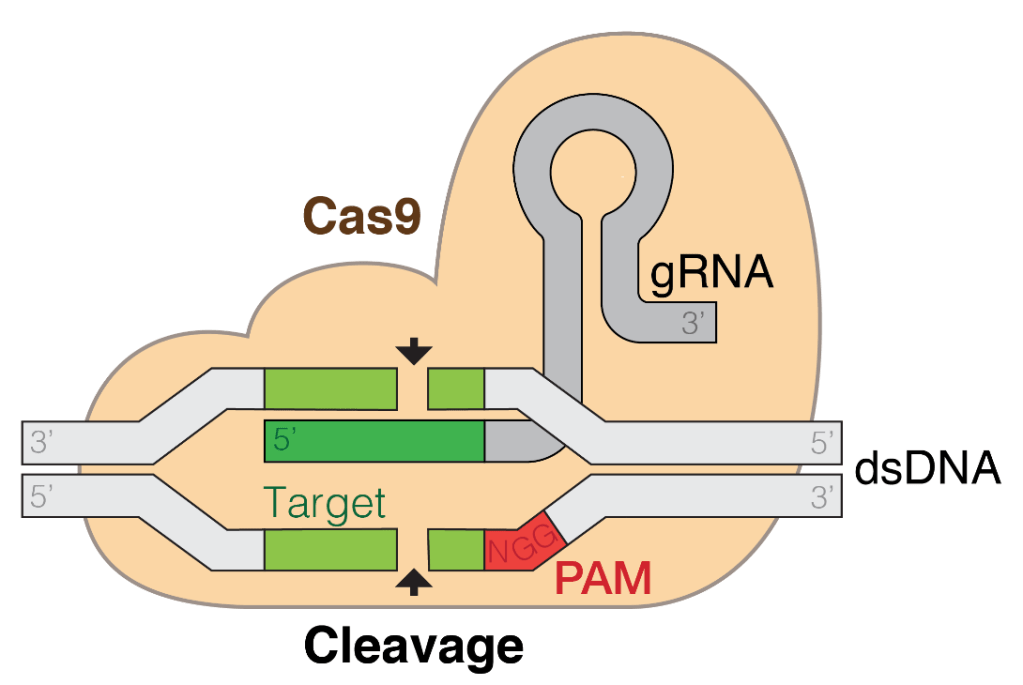
Most biologists would agree that the most extraordinary development of the 21st century has been the discovery and manipulation of the CRISPR/Cas9 method of gene editing – a natural defence mechanism that bacteria use to defend against viral infection which scientists have hijacked to modify the DNA sequence of living cells. The process occurs in nature as bacteria have fragments of viral DNA sequences in their own genomes, which are remnants of previous viral infections. These allow them to recognise and destroy future viral infections, as a form of adaptive immunity. To defend against viruses, the bacteria express the viral DNA fragments as RNAs that become bound by endogenous Cas9 nuclease enzymes. The Cas9 nuclease uses the RNA sequence to guide it to matching regions on infecting viral DNA, which Cas9 then chews up like Pac-Man.

Researchers at the University of California (the labs of Professors Jennifer Doudna and Emmanuelle Charpentier) published a ground-breaking study in Science in 2012, demonstrating how Cas9 could be used to make a double-strand break at any region of DNA if supplied with the right RNA sequence. This revealed the immense potential of CRISPR to correct DNA mutations that cause genetic diseases in humans.
CRISPR is ideal for disrupting or removing genes, and if a donor DNA template sequence is included, it can replace a region, as it is integrated into DNA by the cell’s own “homology-dependent repair” pathway that is activated by the double-strand break. However, perhaps because the classic CRISPR system evolved for viral destruction, it can be rather clumsy when it comes to correcting precise genetic abnormalities, largely inhibiting its uses in research, medicine and agriculture. For example, although CRISPR can remove or replace relatively large regions, it cannot exchange individual DNA bases. The integration of new DNA can also be prone to error (for instance, the new fragment can accidentally be inserted back-to-front). Moreover, accidental binding of the guide strand to the wrong DNA regions leads to off-target cuts, often wreaking havoc across the genome.
Prime editing nicks the limelight
A remarkable new version of the technology was published in Nature in October, which the authors coined “prime editing”. The study was lead-authored by Post-Doctoral Fellow Andrew V. Anzalone, from Professor David Liu’s laboratory at Harvard University.
Prime editing is an ingenious improvement on traditional CRISPR that uses an impaired Cas9 nuclease that cuts only one strand of the DNA. It also does not require a donor DNA template for an insertion. Instead the prime editing guide RNA (pegRNA) is lengthened and encodes the replacement sequence, which is copied by a reverse transcriptase enzyme that has been conjugated to the Cas9. To correct a mutation, the Cas9 “nicks” one strand of the DNA and the pegRNA becomes fused to it, acting as a template for the reverse transcriptase enzyme to copy, which produces a double-stranded sequence that is inserted into the DNA.
Professor Lui’s group previously developed a novel method of individual base editing, which can correct transition mutations (A to G or C to T, since the structure of the bases are similar) but cannot correct transversion mutations (G to T, or A to C). The group have now used prime editing successfully to correct the mutation causing sickle cell disease (an T•A-to-A•T transversion mutation) in blood cells – a mutation that cannot be repaired by CRISPR nor base editing. Unexpectedly, prime editing also caused far fewer off-target cuts compared to traditional CRISPR, making it a much safer system.
Overall, prime editing removes the need for a risky double-strand break, is suitable for all individual base changes, and it is less prone to mistakes like inverted sequences. There are many more obstacles to overcome before prime editing could become part of routine patient treatment, such as improving its delivery into human tissues, however this fascinating study represents a major step in the honing of CRISPR technologies.
Regulatory and policy issues in human gene editing
Last year a scientist appalled the world as he presented baby twin girls whose embryo he had modified in vitro using CRISPR – the technology is far from being safe enough for use in humans. The twins are currently believed to be healthy (although their futures remain uncertain as more of their genes will be required during development). His study highlighted the need for clear international guidelines and open, evidence-based public discourse on genetic modification, which do not currently exist. Central to this discussion will be distinguishing between germline gene editing (where the mutation will be inherited by all future cells in the individual, such as in an embryo) and somatic gene editing (for instance, correcting faulty gene expression in a developed tissue such as a tumour, which will not affect future offspring).

Genetic selection (in the form of breeding) has been around for centuries – but to what extent are CRISPR technologies different from traditional methods of genetic modification? The advent of CRISPR made editing the human genome uncomfortably close on the horizon, bringing an enticing promise of curing many genetic diseases yet disturbing questions regarding eugenics. In response, many countries are in the process of reassessing their national regulatory frameworks, although progress is happening far slower than research in the billion-dollar field. Key questions remain – if CRISPR is used to create a version of a plant that is identical from one found in nature (rather than something abnormal, for example, a rice plant modified to include a gene producing vitamin A), then should it be treated as a GMO, as the Court of Justice of the European Union (ECJ) concluded in 2018? What sort of effects could it have on biodiversity and the environment? At what point should a CRISPR-based method be classed as safe enough for use in humans – what sort of error-rate will be acceptable? No medicine is 100% “safe” and there are risks with every medical intervention, although the permanence of a mistake generated by a CRISPR is frightening. These important questions require communication between many disciplines of science as well as lawyers and the general population, and must be fast addressed.
About the author
Zoe Angel is a Research Fellow at the Centre for Cancer Research and Cell Biology, Queen’s University Belfast.
Further reading list
Anzalone, AV, Randolph, PB, Davis, JR. (2019). Search-and-replace genome editing without double-strand breaks or donor DNA. Nature. 576 (7785), 149–157.
Bhaya, D., Davison, M., Barrangou, R.. (2011). CRISPR-Cas Systems in Bacteria and Archaea: Versatile Small RNAs for Adaptive Defense and Regulation. Annual Review of Genetics. 45 (1), 273-297.
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E. (2012). A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science. 17 (337), 816-21.
Judgement of the Court (Grand Chamber), 25 July 2018 in Case C-528/16
Komor AC, Kim YB, Packer MS, Zuris JA, Liu DR.. (2016). Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage.. Nature. 19 (533), 420-4.
Kosicki M, Tomberg K, Bradley A.. (2018). Repair of double-strand breaks induced by CRISPR– Cas9 leads to large deletions and complex rearrangements. Nature Biotechnology. 36 (8), 765-771.

