By Kimberley D. Morrison, University of Dundee
Background
Affecting approximately 20-30% of children and 2-3% of adults globally, eczema (which is synonymous with atopic dermatitis) is the most common skin disease at present. Primarily characterised as an itchy and inflammatory skin disease, research has aimed to answer the question of what causes eczema? The answer, as expected, is at present incomplete and complex. Eczema is a disease caused by a multitude of factors. These range from internal factors, such as genetic differences – which results in alterations at the molecular level – and our immune response to stimuli, to external factors i.e. the environment in which we live. This entry briefly, and simply, summarises what we understand specifically about the genetics of eczema to date, and how we can use sophisticated methods across research disciplines to further study this complex and troublesome skin disease.
Introducing the link between genetics and eczema
In simplest terms, the skin is formed from two main components: the dermis (mainly composed of cells named fibroblasts) and the epidermis (mainly composed of multiple layers of cells named keratinocytes, which are at various stages of differentiation) (fig. 1). Your skin is incredibly important in providing a protective barrier. When functioning correctly, it keeps everything that should remain external, out. Conversely, it keeps everything that should remain internal, in. Additionally, your skin provides structure and helps control water loss etc. (fig. 1). Therefore, as you can imagine, when this barrier becomes defective problems can arise; such as the development of eczematous skin. But how do these problems arise?

Figure 1. A summary of eczema. The skin is composed primarily of two components, the dermis and epidermis. The dermis contains the fibroblasts. The epidermis contains cell layers (from the stratum basale to the stratum corneum) which are composed of keratinocytes undergoing terminal differentiation as they progress through these layers. When the skin barrier is functioning normally it provides a barrier preventing the entry or release of various things. Conversely, when the skin barrier is impaired it becomes ‘leaky’ and does not function normally, increasing the risk of developing eczema (atopic dermatitis). This barrier can become leaky and impaired through environmental exposures, the immune response and how our skin is made i.e. genetics.
As alluded to previously, your genetics play a big role in increasing the risk of the development of eczema. Many studies have been conducted which show eczema to be a highly heritable disease – identical twins have an 80% chance of developing eczema if their sibling suffers from it compared to a 20% chance for non-identical twins. With this knowledge, research, through Genome-Wide Association Studies (GWAS), has identified a growing number of risk loci (specific chromosome position) associated with eczema. While no one gene has been identified in causing eczema alone, to date, the gene that has the biggest association with eczema is FLG, with an odds ratio of around 3 (i.e. possessing a loss-of-function variant in FLG can increase your risk of developing eczema 300%).
FLG encodes filaggrin (filament-aggregating protein), a protein important in various processes involved in skin barrier formation and development. This association was discovered in 2006 and since a number of loss-of-function mutations have been identified for FLG. Additionally, a number of variants in other genes have been named as potentially important in determining eczema development; more than 30 risk loci have now been identified – with further research required to confirm the degree of importance for each.
What next?
While the identification of these risk loci is invaluable to furthering our understanding of the genetic risks associated with eczema, it is imperative we find the molecular consequences. With an increased understanding we can shape the future of treatment options for eczema, with the aim to improve upon current therapies. One way we can study eczema in the lab is through the use of organotypic modelling. We know that eczema is a multifaceted (many sided) disease caused by many things working together. This can make research into the genetic basis of the disease somewhat challenging.
This is where organotypic modelling is useful because not only are the external conditions controlled, it is a simplified model whereby the only cells present are the fibroblasts (forming the dermis) and the keratinocytes (forming the epidermis). Using this model, we can target ‘suspect’ genes initially through knock-down experiments and perform various experiments that allow us to assess skin barrier function and health. This can be assessed in a number of ways from functional testing – which allows us to determine how dry or ‘leaky’ the barrier is, indicative of eczema – to histology, allowing us to look for phenotypic differences. Larger scale, we can look for changes within the global-proteome using detailed proteomic analyses. Combined, these experiments allow us to confidently piece together and make sense of, what is, a complex disease.
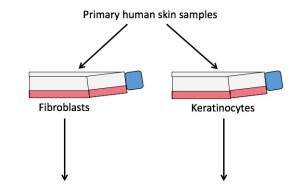
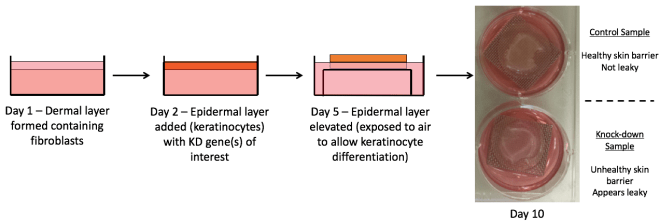
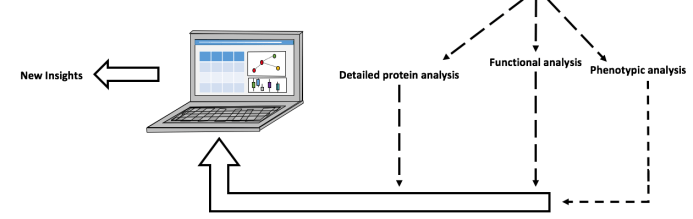
Figure 2. Schematic diagram of researching eczema through organotypic modelling. Primary fibroblasts and keratinocytes from donor human skin samples are separated and cultured. The dermal layer is made up on day 1 from the primary fibroblasts – these are contained within a fibrin gel matrix. Using reverse transfection, gene(s) of interest are knocked down (KD) in the primary keratinocytes, which are added on top of the dermal layer to form the epidermis. On day 5 the model gel is lifted onto a grid so that the epidermis is exposed to the air allowing for keratinocyte differentiation to take place. On day 10 the organotypics are collected for various analyses to assess impact on skin architecture and molecular processes.
The first genetic studies conducted over 10 years ago have shaped our understanding of eczema – however, it is clear more work is required. Using sophisticated methods in parallel – such as organotypic modelling, functional and phenotypic assessment and detailed proteomic analyses – we can continue to fill in the gaps and use such knowledge to shape the development of treatment in the future. Altogether, I aim to use the above methods in conjunction with previously published knowledge gained in previous genetic studies throughout my PhD. This will allow me to identify proteins implicated in eczema helping us uncover previously unknown aspects of the pathology.
Further reading:
- Hammers C.M., Tang H, Chen J, Emtenani S, Zheng Q & Stanley J.R. Research Techniques Made Simple: Mass Spectrometry for Analysis of Proteins in Dermatological Research. Journal of Investigative Dermatology 138: 1236-1242 (2018)
- Brown S.J. Molecular mechanisms in atopic eczema: insights gained from genetic studies. Journal of Pathology 241: 140-145 (2017)
- Palmer C.N. et al. Common loss-of-function variants of the epidermal barrier protein filaggrin are a major predisposing factor for atopic dermatitis. Nature Genetics 38(4): 441-446 (2006)
About me:

I am a first year PhD student within the School of Medicine at the University of Dundee. Here I am studying the genetic mechanisms in skin differentiation and barrier function. My main aim is to further our understanding into the genetic basis and molecular mechanisms of eczema.
Previously I obtained a BSc (Hons) in Cell Biology from the University of St. Andrews and an MRes in Cancer Biology from the University of Dundee.

